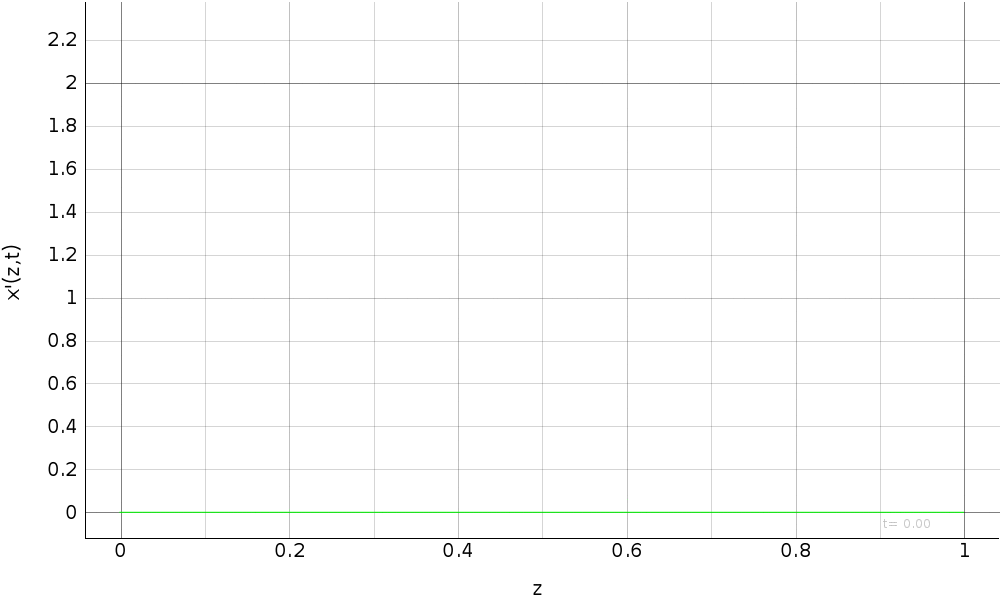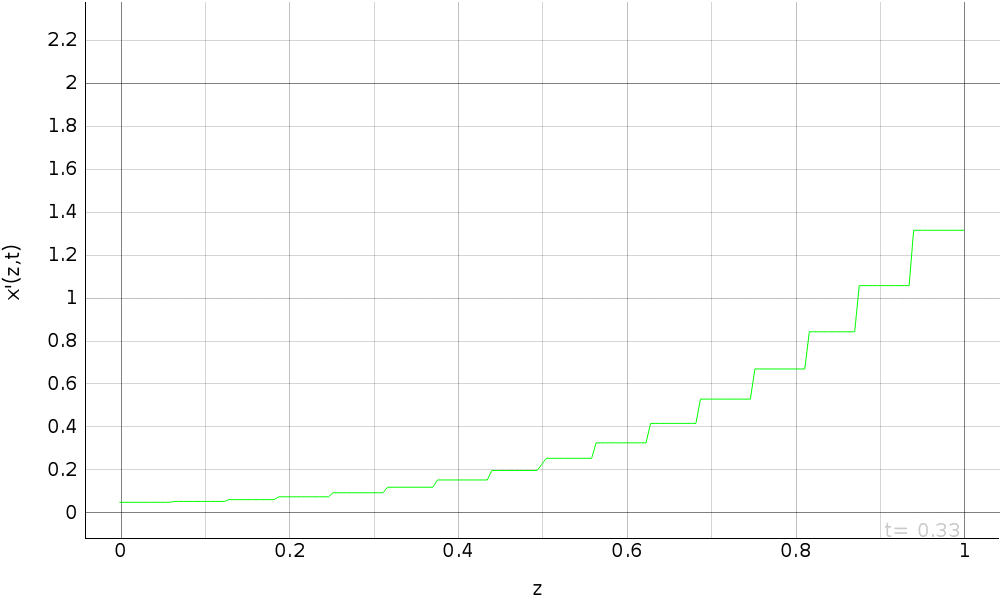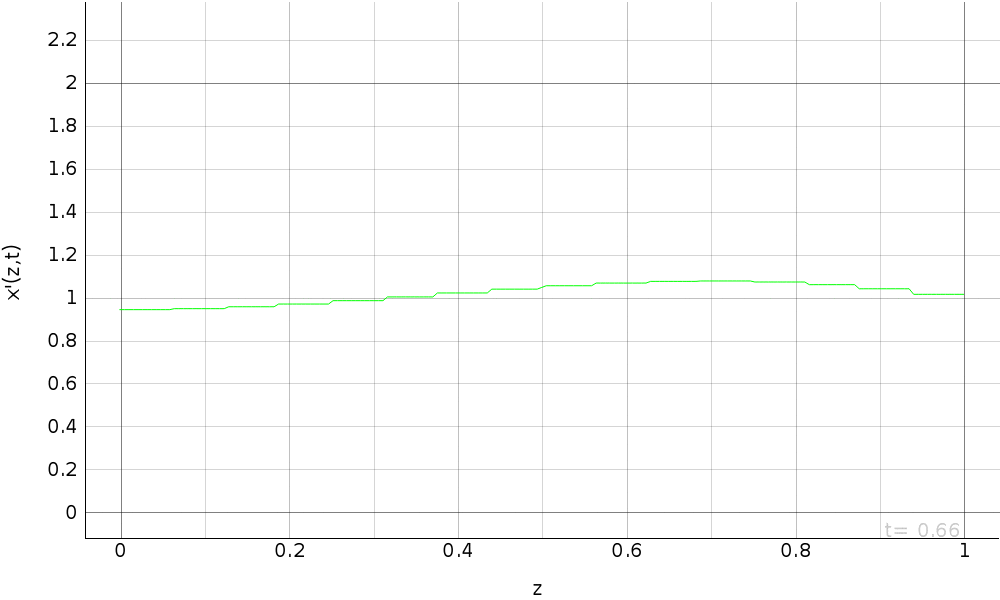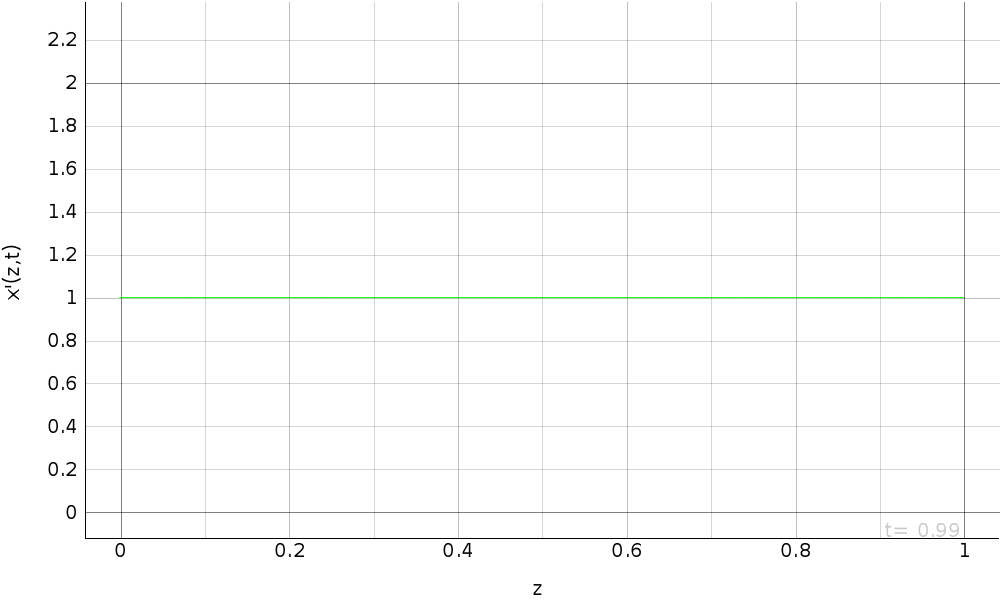
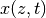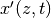R.a.d. eq. with dirichlet b.c. (fem approximation)¶
Simulation of the reaction-advection-diffusion equation with dirichlet boundary
condition by 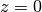 and dirichlet actuation by
and dirichlet actuation by 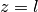 .
.
![\begin{align*}
\dot{x}(z,t) = a_2x''&(z,t) + a_1x'(z,t) + a_0x(z,t) && z\in (0, l), t>0\\
x(z,0) &= x_0(z) && z\in [0,l]\\
x(0,t) &= 0 && t>0\\
x(l,t) &= u(t) && t>0
\end{align*}](../_images/math/7a1bc4756a0f858ea28dfb25fc778ee32d9ab551.png)
example: heat equation
|
|
|
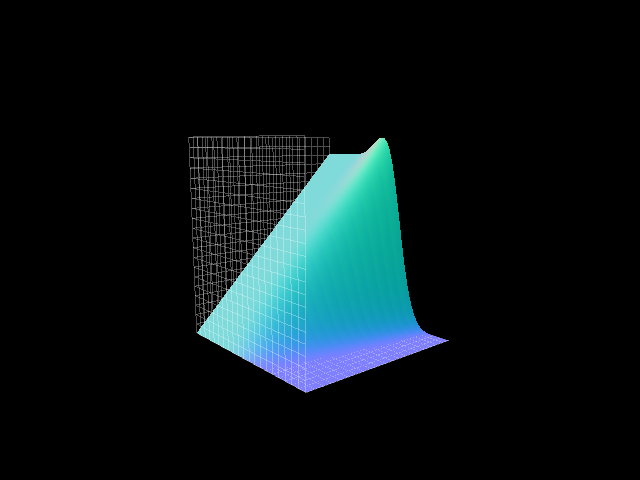
|
with:
inital functions
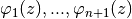
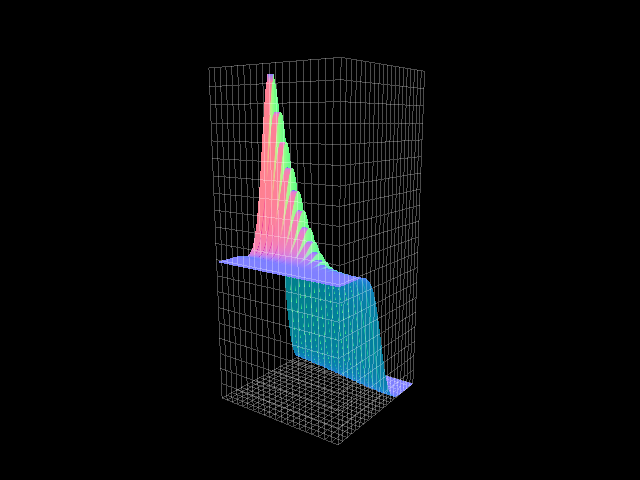
test functions
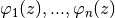

where the functions
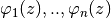 met the homogeneous b.c.
met the homogeneous b.c.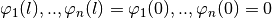
only
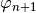 can draw the actuation
can draw the actuationfunctions
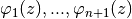 e.g. from type
e.g. from type pyinduct.shapefunctions.LagrangeFirstOrderorpyinduct.shapefunctions.LagrangeSecondOrder, seepyinduct.shapefunctions
approach:
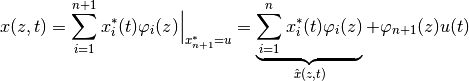
weak formulation…
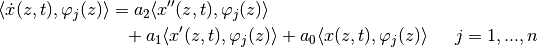
… and derivation shift to work with lagrange 1st order initial functions
![\begin{align*}
\langle\dot{x}(z,t),\varphi_j(z)\rangle &=
\overbrace{[a_2 [x'(z,t)\varphi_j(z)]_0^l}^{=0} - a_2 \langle x'(z,t),\varphi'_j(z)\rangle \\
&\hphantom =+
a_1 \langle x'(z,t), \varphi_j(z)\rangle +
a_0 \langle x(z,t), \varphi_j(z)\rangle && j=1,...,n \\
\langle\dot{\hat{x}}(z,t),\varphi_j(z)\rangle + \langle\varphi_{N+1}(z),\varphi_j(z)\rangle \dot u(t) &= - a_2 \langle \hat x'(z,t),\varphi'_j(z)\rangle - a_2 \langle \varphi'_{N+1}(z),\varphi'_j(z)\rangle u(t) \\
&\hphantom =+
a_1 \langle \hat x'(z,t), \varphi_j(z)\rangle + a_1 \langle \varphi'_{N+1}(z), \varphi_j(z)\rangle u(t) + \\
&\hphantom =+
a_0 \langle \hat x(z,t), \varphi_j(z)\rangle + a_0 \langle \varphi_{N+1}(z), \varphi_j(z)\rangle u(t) && j=1,...,n
\end{align*}](../_images/math/0d4430d18e333b3aa57a96c546510af6f573499d.png)
leads to state space model for the weights
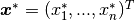 :
:
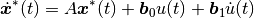
input derivative elimination through the transformation:
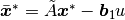
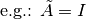
leads to
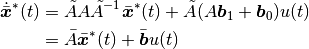
source code:
import numpy as np
import pyinduct as pi
import pyinduct.parabolic as parabolic
def run(show_plots):
n_fem = 17
T = 1
l = 1
y0 = -1
y1 = 4
param = [1, 0, 0, None, None]
# or try these:
# param = [1, -0.5, -8, None, None] # :)))
a2, a1, a0, _, _ = param
temp_domain = pi.Domain(bounds=(0, T), num=100)
spat_domain = pi.Domain(bounds=(0, l), num=n_fem * 11)
# initial and test functions
nodes = pi.Domain(spat_domain.bounds, num=n_fem)
fem_base = pi.LagrangeFirstOrder.cure_interval(nodes)
act_fem_base = pi.Base(fem_base[-1])
not_act_fem_base = pi.Base(fem_base[1:-1])
vis_fems_base = pi.Base(fem_base)
pi.register_base("act_base", act_fem_base)
pi.register_base("sim_base", not_act_fem_base)
pi.register_base("vis_base", vis_fems_base)
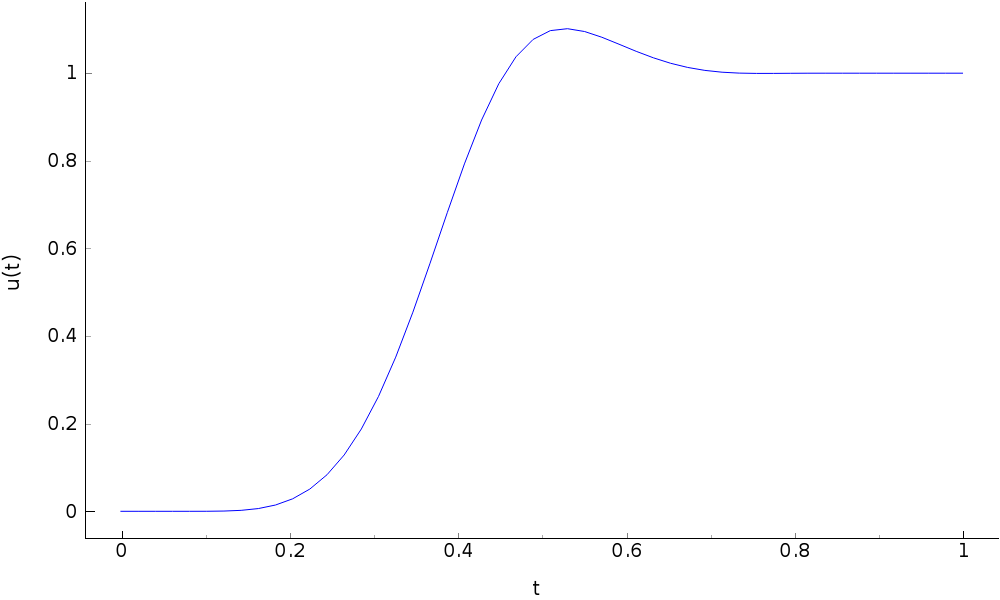
# trajectory
u = parabolic.RadFeedForward(l, T,
param_original=param,
bound_cond_type="dirichlet",
actuation_type="dirichlet",
y_start=y0, y_end=y1)
# weak form
x = pi.FieldVariable("sim_base")
x_dt = x.derive(temp_order=1)
x_dz = x.derive(spat_order=1)
phi = pi.TestFunction("sim_base")
phi_dz = phi.derive(1)
act_phi = pi.ScalarFunction("act_base")
act_phi_dz = act_phi.derive(1)
weak_form = pi.WeakFormulation([
# ... of the homogeneous part of the system
pi.IntegralTerm(pi.Product(x_dt, phi),
limits=spat_domain.bounds),
pi.IntegralTerm(pi.Product(x_dz, phi_dz),
limits=spat_domain.bounds,
scale=a2),
pi.IntegralTerm(pi.Product(x_dz, phi),
limits=spat_domain.bounds,
scale=-a1),
pi.IntegralTerm(pi.Product(x, phi),
limits=spat_domain.bounds,
scale=-a0),
# ... of the inhomogeneous part of the system
pi.IntegralTerm(pi.Product(pi.Product(act_phi, phi),
pi.Input(u, order=1)),
limits=spat_domain.bounds),
pi.IntegralTerm(pi.Product(pi.Product(act_phi_dz, phi_dz),
pi.Input(u)),
limits=spat_domain.bounds,
scale=a2),
pi.IntegralTerm(pi.Product(pi.Product(act_phi_dz, phi),
pi.Input(u)),
limits=spat_domain.bounds,
scale=-a1),
pi.IntegralTerm(pi.Product(pi.Product(act_phi, phi),
pi.Input(u)),
limits=spat_domain.bounds,
scale=-a0)],
name="main_system")
# system matrices \dot x = A x + b0 u + b1 \dot u
cf = pi.parse_weak_formulation(weak_form)
ss = pi.create_state_space(cf)
a_mat = ss.A[1]
b0 = ss.B[0][1]
b1 = ss.B[1][1]
# transformation into \dot \bar x = \bar A \bar x + \bar b u
a_tilde = np.diag(np.ones(a_mat.shape[0]), 0)
a_tilde_inv = np.linalg.inv(a_tilde)
a_bar = (a_tilde @ a_mat) @ a_tilde_inv
b_bar = a_tilde @ (a_mat @ b1) + b0
# simulation
def x0(z):
return 0 + y0 * z
start_func = pi.Function(x0, domain=spat_domain.bounds)
full_start_state = np.array([pi.project_on_base(start_func,
pi.get_base("vis_base")
)]).flatten()
initial_state = full_start_state[1:-1]
start_state_bar = a_tilde @ initial_state - (b1 * u(time=0)).flatten()
ss = pi.StateSpace(a_bar, b_bar, base_lbl="sim", input_handle=u)
sim_temp_domain, sim_weights_bar = pi.simulate_state_space(ss,
start_state_bar,
temp_domain)
# back-transformation
u_vec = np.reshape(u.get_results(sim_temp_domain), (len(temp_domain), 1))
sim_weights = sim_weights_bar @ a_tilde_inv + u_vec @ b1.T
# visualisation
plots = list()
save_pics = False
vis_weights = np.hstack((np.zeros_like(u_vec), sim_weights, u_vec))
eval_d = pi.evaluate_approximation("vis_base",
vis_weights,
sim_temp_domain,
spat_domain,
spat_order=0)
der_eval_d = pi.evaluate_approximation("vis_base",
vis_weights,
sim_temp_domain,
spat_domain,
spat_order=1)
if show_plots:
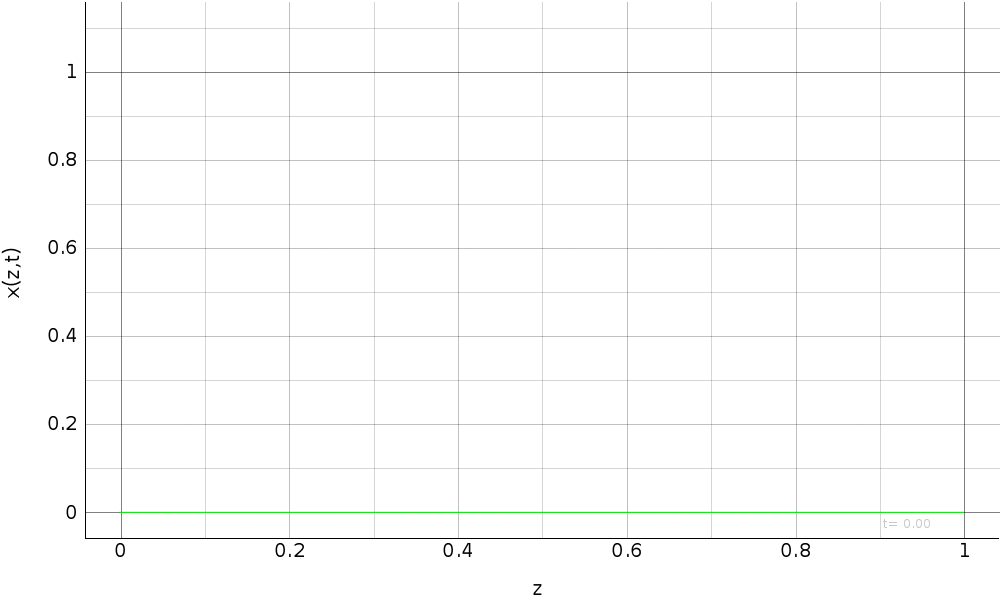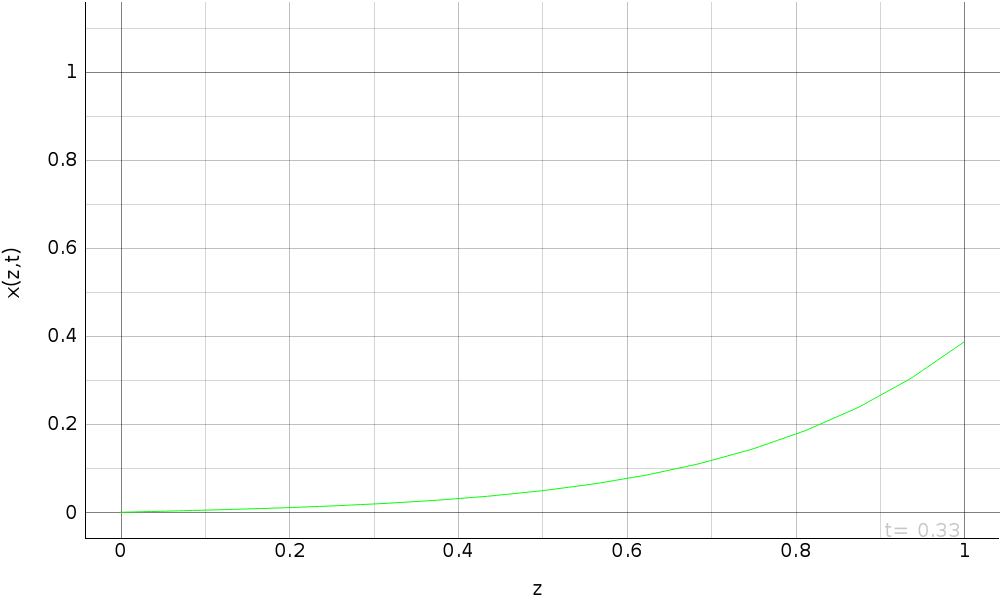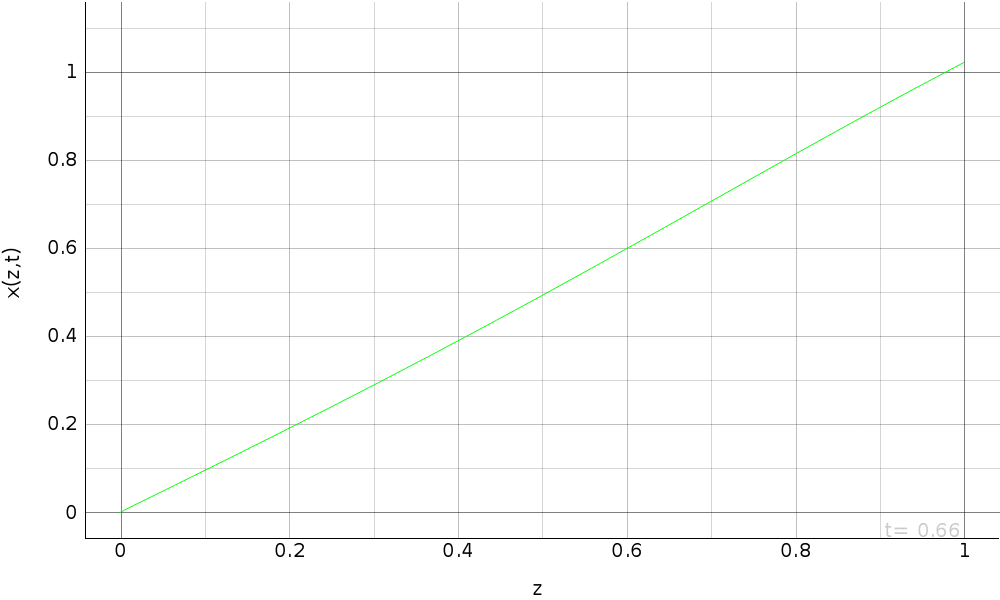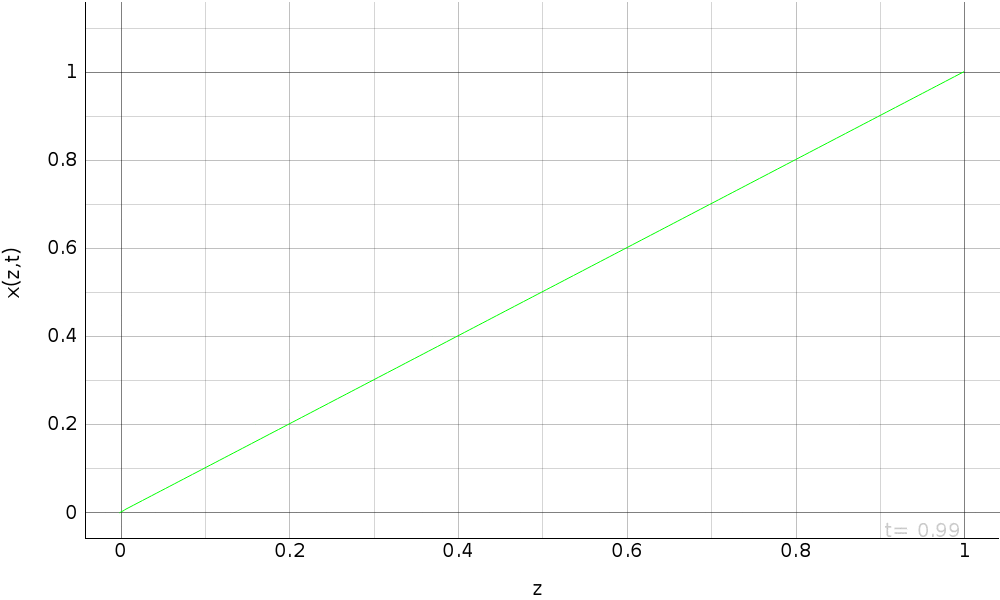
plots.append(pi.PgAnimatedPlot(eval_d,
labels=dict(left='x(z,t)', bottom='z'),
save_pics=save_pics))
plots.append(pi.PgAnimatedPlot(der_eval_d,
labels=dict(left='x\'(z,t)', bottom='z'),
save_pics=save_pics))
win1 = pi.surface_plot(eval_d, title="x(z,t)")
win2 = pi.surface_plot(der_eval_d, title="x'(z,t)")
# save pics
if save_pics:
path = pi.save_2d_pg_plot(u.get_plot(), 'rad_dirichlet_traj')[1]
win1.gl_widget.grabFrameBuffer().save(path + 'rad_dirichlet_3d_x.png')
win2.gl_widget.grabFrameBuffer().save(path + 'rad_dirichlet_3d_dx.png')
pi.show()
pi.tear_down(("act_base", "sim_base", "vis_base"))
if __name__ == "__main__":
run(True)
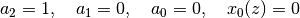
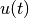 –>
–>